Inference of differential kinase interaction networks with KINference is published in Bioinformatics (https://doi.org/10.1093/bioinformatics/btaf349). KINference combines computation of a baseline kinase interaction network (KIN) representing the space of all possible kinase–substrate links with filters applied to nodes and edges to identify differentially active subnetworks that are relevant in the context of a specific phosphoproteomics dataset. For the node filters, we rely on functional relevance and differential phosphorylation scores; for the edge filters, we make use of prize-collecting Steiner trees and correlations between phosphorylation sites of kinases and their target proteins. KINference is available as an R package at https://github.com/bionetslab/KINference.
This work is part of WP2: context-specific PPI networks.
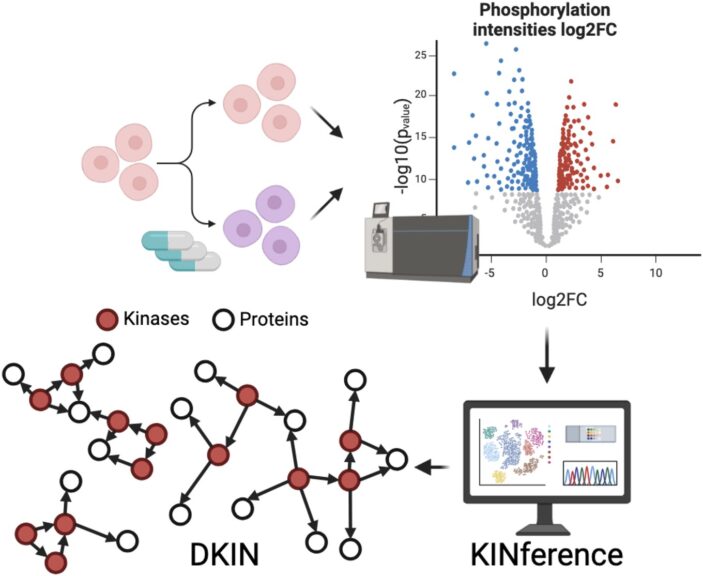
Inference of differential kinase interaction networks with KINference is published in Bioinformatics (https://doi.org/10.1093/bioinformatics/btaf349). KINference combines computation of a baseline kinase interaction network (KIN) representing the space of all possible kinase–substrate links with filters applied to nodes and edges to identify differentially active subnetworks that are relevant in the context of a specific phosphoproteomics dataset. For the node filters, we rely on functional relevance and differential phosphorylation scores; for the edge filters, we make use of prize-collecting Steiner trees and correlations between phosphorylation sites of kinases and their target proteins. KINference is available as an R package at https://github.com/bionetslab/KINference.
This work is part of WP2: context-specific PPI networks.